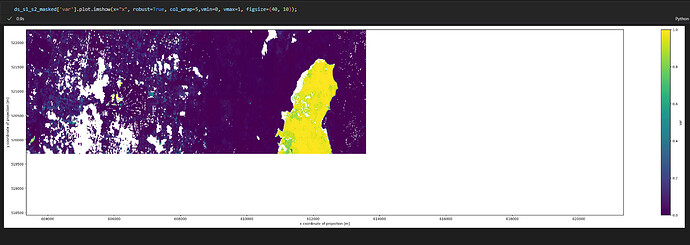
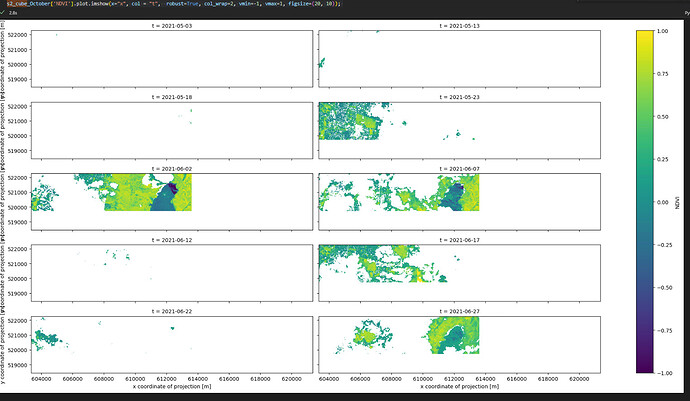
Hi Andrea,
I tried it, in fact, due to no longer loading SCL and CLP in the first cube, we need to adjust the indices.
I also made a minor adjustment to masking with SCL and CLP. This example now works here:
from openeo.processes import exp,log,if_
connection = openeo.connect("openeo-dev.vito.be").authenticate_oidc()
start_date = '2021-06-01'
spatial_extent = {'west': -74.06810760, 'east': -73.90597343, 'south': 4.689864510, 'north': 4.724080996,
'crs': 'epsg:4326'} # colombia
zone = "tropical"
start_date_dt_object = datetime.strptime(start_date, '%Y-%m-%d')
end_date = (start_date_dt_object + relativedelta(months=+1)).date() ## End date, 1 month later (1st Feb. 2021)
start_date_exclusion = (start_date_dt_object + relativedelta(months=-1)).date()
LOOKUPTABLE = {
"tropical": {
"S1": lambda vv: 1 / (1 + exp(- (-7.17 + (-0.48 * vv)))),
"S2": lambda ndvi, ndwi: 1 / (1 + exp(- (0.845 + (2.14 * ndvi) + (13.5 * ndwi)))),
"S1_S2": lambda vv, ndwi: 1 / (1 + exp(- (-2.64 + (-0.23 * vv) + (8.6 * ndwi)))),
},
"subtropical":
{
"S1": lambda vv, vh: 1 / (1 + exp(- (-8.1 + (-0.13 * vv) + (-0.27 * vh)))),
"S2": lambda ndvi, ndwi: 1 / (1 + exp(- (0.845 + (2.14 * ndvi) + (13.5 * ndwi)))),
"S1_S2": lambda vv, ndwi: 1 / (1 + exp(- (-2.64 + (-0.23 * vv) + (8.6 * ndwi)))),
}
}
s2_cube = connection.load_collection(
'SENTINEL2_L2A_SENTINELHUB',
spatial_extent=spatial_extent,
temporal_extent=[start_date_exclusion, end_date],
bands=['B02', 'B03', 'B04', 'B08', 'sunAzimuthAngles', 'sunZenithAngles'])
# This will avoid loading data from sentinelhub which is then discarded later on
s2_cube_masking = connection.load_collection(
'SENTINEL2_L2A_SENTINELHUB',
spatial_extent=spatial_extent,
temporal_extent=[start_date_exclusion, end_date],
bands=['CLP', 'SCL'])
# Scene classification data, based on Sen2Cor
# scl == 3 Cloud Shadows
# scl == 8 Clouds medium probability
# scl == 9 Clouds high probability
# scl == 10 Cirrus
# scl == 11 Snow / Ice
scl = s2_cube_masking.band("SCL")
mask_scl = (scl == 3) | (scl == 8) | (scl == 9) | (scl == 10) | (scl == 11)
# mask {1 = cloud, 0 = free cloud}
#s2_cube = s2_cube.mask(mask_scl)
# CLP (cloud probabilities) based on S2cloudless (160 blocks and 10m resolution)
clp = s2_cube_masking.band("CLP")
mask_clp = mask_scl | (clp / 255) > 0.3
# mask {1 = cloud, 0 = free cloud}
s2_cube = s2_cube.mask(mask_clp)
s2_count = s2_cube.filter_bands(bands=["B08"])
s2_count = s2_count.reduce_dimension(reducer=lambda data: data.count(), dimension="t")
s2_count = s2_count.rename_labels("bands", ["count"])
s2_cube = append_indices(s2_cube, ["NDWI","NDVI"]) ## index 6,7
#s2_cube = append_index(s2_cube, "NDVI") ## index 8
def water_function(data):
return LOOKUPTABLE[zone]["S2"](ndwi=data[6], ndvi=data[7])
s2_cube_water = s2_cube.reduce_dimension(reducer=water_function, dimension="bands")
s2_cube_water = s2_cube_water.add_dimension("bands", "water_prob", type="bands")
s2_cube_water_threshold = s2_cube_water.apply_dimension(dimension="bands", process=lambda x: if_(x > 0.75, x, 0))
s2_cube_water_threshold = s2_cube_water_threshold.rename_labels("bands", ["w_T75"])
s2_cube_water_sum = s2_cube_water_threshold.reduce_dimension(reducer="sum", dimension="t")
s2_cube_water_sum = s2_cube_water_sum.rename_labels("bands", ["sum"])
s2_cube_swf = s2_cube_water_sum.resample_cube_spatial(s2_count) / s2_count
s2_cube_swf = s2_cube_swf.rename_labels("bands", ["swf"])
## calculate the median s2 water probability for 1-month.
s2_median_water = s2_cube_water.filter_temporal([start_date, end_date]).median_time()
s2_cube_median = s2_cube.filter_temporal([start_date, end_date]).median_time()
## Get Sentinel-1 data for a 1 month window and convert to ARD data.
s1_cube = connection.load_collection(
'SENTINEL1_GRD',
spatial_extent=spatial_extent,
temporal_extent=[start_date, end_date],
bands=['VH', 'VV'],
properties={"polarization": lambda p: p == "DV"})
# s1_cube = s1_cube.ard_normalized_radar_backscatter(elevation_model="COPERNICUS_30")
s1_cube = s1_cube.sar_backscatter(coefficient="gamma0-terrain", mask=True, elevation_model="COPERNICUS_30")
s1_cube = s1_cube.rename_labels("bands", ["VH", "VV", "mask", "incidence_angle"])
s1_cube_mask = s1_cube.band("mask")
# radar shadow mask has value 2
s1_mask_RS = (s1_cube_mask == 2)
s1_cube = s1_cube.mask(s1_mask_RS)
def log_(x):
return 10 * log(x, 10)
s1_median = s1_cube.median_time().apply(log_)
def s1_water_function(data):
return LOOKUPTABLE[zone]["S1"](vv=data[1])
s1_median_water = s1_median.reduce_dimension(reducer=s1_water_function, dimension="bands")
exclusion_mask = (s1_median_water.resample_cube_spatial(s2_cube_swf) > 0.5) & (s2_cube_swf < 0.33)
exclusion_mask = exclusion_mask * 1.0
#s1_median_water_mask = s1_median_water.mask(exclusion_mask.resample_cube_spatial(s1_median_water))
#s1_median_water_mask = s1_median_water_mask * 1.0
def s1_s2_water_function(data):
return LOOKUPTABLE[zone]["S1_S2"](vv=data[0], ndwi=data[1])
## Get S1 VV, and S2 NDWI, and apply s1_s2_water function
s1_s2_cube = s1_median.filter_bands(['VV']).resample_cube_spatial(s2_cube_median).merge_cubes(
s2_cube_median.filter_bands(['NDWI'])) # works
s1_s2_water = s1_s2_cube.reduce_dimension(reducer=s1_s2_water_function, dimension="bands").add_dimension("bands",
"var",
type="bands")
s1_s2_water = s1_s2_water.rename_labels("bands", ["var"])
s1_s2_water = s1_s2_water.filter_bands(bands=['var'])
exclusion_mask = exclusion_mask.add_dimension("bands", "var", type="bands")
exclusion_mask = exclusion_mask.rename_labels("bands", ["var"])
exclusion_mask = exclusion_mask.filter_bands(bands=['var'])
masked = s1_s2_water.mask(exclusion_mask.resample_cube_spatial(s1_s2_water))
s1_s2_masked_save = masked.save_result(format='netCDF') # GTiff #netCDF
my_job = s1_s2_masked_save.send_job(title="s1_s2_masked_save")
results = my_job.start_and_wait().get_results()
results.download_files('s1_s2_masked_save')