Hey all,
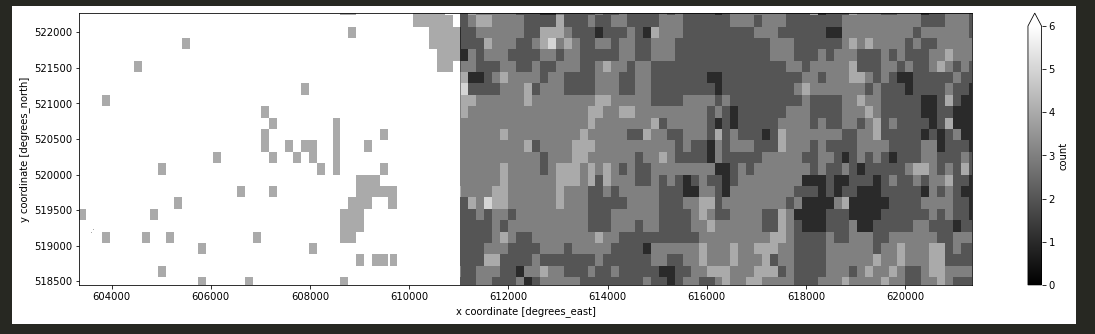
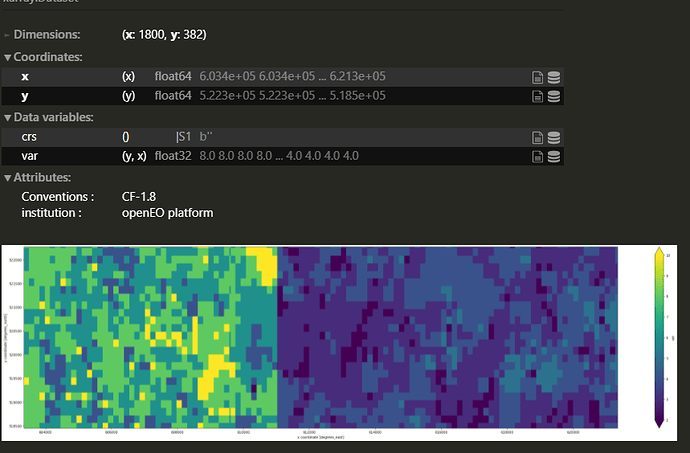
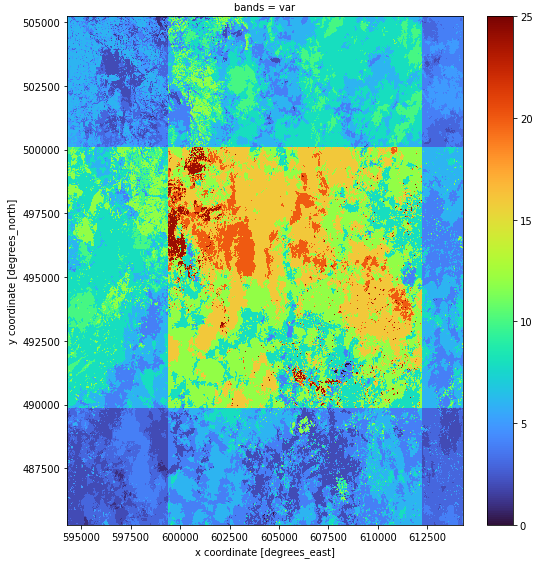
Please can you advice, why after applying reduce_dimension(reducer = "count", dimension = "t") , the ouput data shows an incorect values in final raster, mainly in the left side of image?
I aim to reduce the dimension of a datacube by calculating the number of images with a valid no-mask value at each pixel across the stack of B02 band, e.g., in this case we have 6 scenes so the final raster should have the max value equal 6 but many scenes are masked so pixels should reach values below 6.
## Get the Sentinel-2 data for a 3 month window.
start_date = '2021-01-01'
end_date = '2021-01-31'
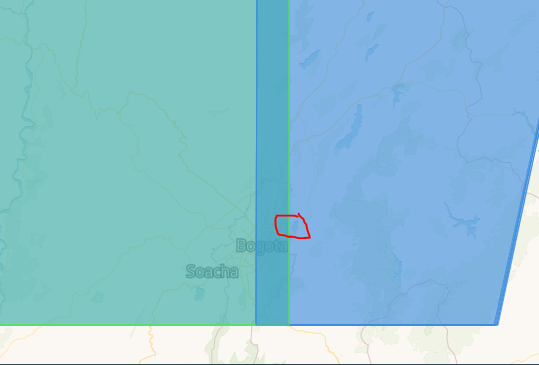
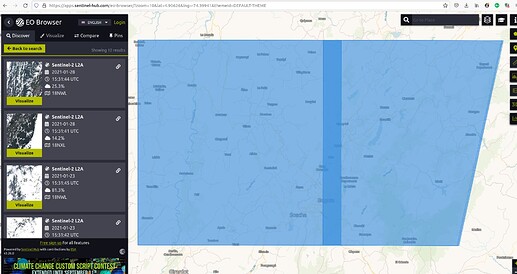
spatial_extent = {'west': -74.06810760, 'east': -73.90597343, 'south': 4.689864510, 'north': 4.724080996, 'crs': 'epsg:4326'}
s2_cube = connection.load_collection(
'SENTINEL2_L2A_SENTINELHUB',
spatial_extent = spatial_extent,
temporal_extent = [start_date, end_date],
bands = ['B02', 'B03', 'B04', 'B08', 'CLP', 'SCL', 'sunAzimuthAngles'] )
clp = s2_cube.band("CLP")
mask = (clp / 255) > 0.3
s2_cube = s2_cube.mask(mask)
ds_s2_cube_v1 = xr.open_dataset("s2_cube_v4.nc").load()
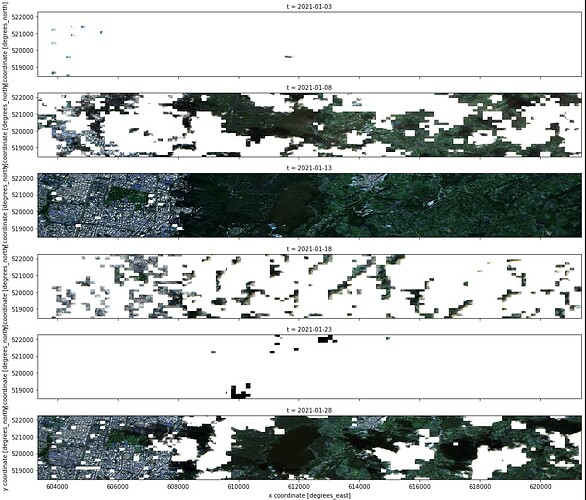
ds_s2_cube_v1[['B02', 'B03', 'B04']].to_array().plot.imshow(x="x", robust=True, col ="t", col_wrap=1,vmin=0, vmax=3000, figsize=(14, 6));
s2_count = s2_cube.filter_bands(bands = ["B02"]).reduce_dimension(reducer = "count", dimension = "t")
s2_count = s2_count.add_dimension("bands", "count", type = "bands")
ds_s2_cube_count_v2 = xr.open_dataset("s2_count_V4.nc").load()
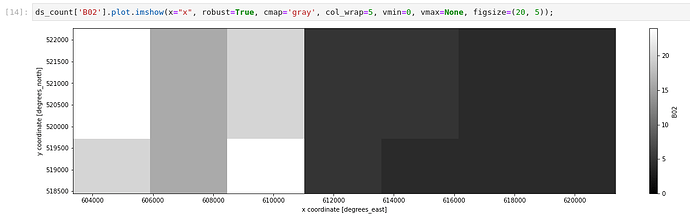
ds_s2_cube_count_v2['count'].plot.imshow(x="x", robust=True, cmap='gray', col_wrap=5,vmin=0, vmax=6, figsize=(20, 5));
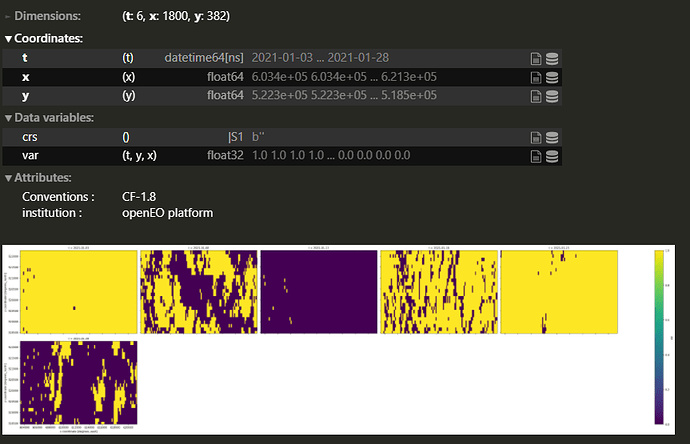
S2 RGB scenes after cloud masking look like these:
The final output:
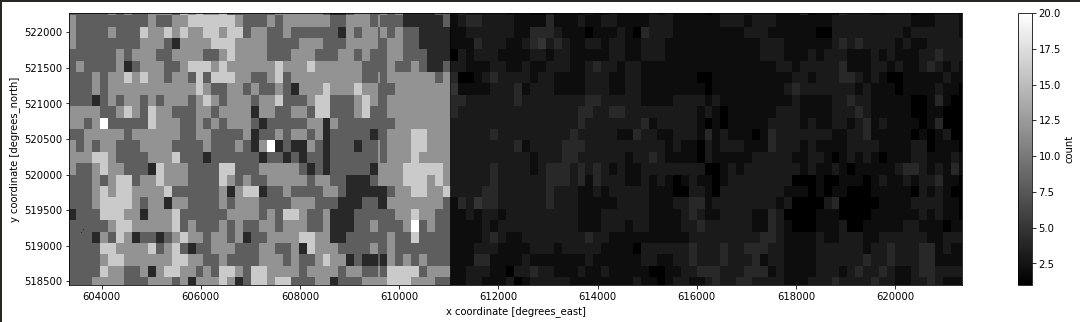
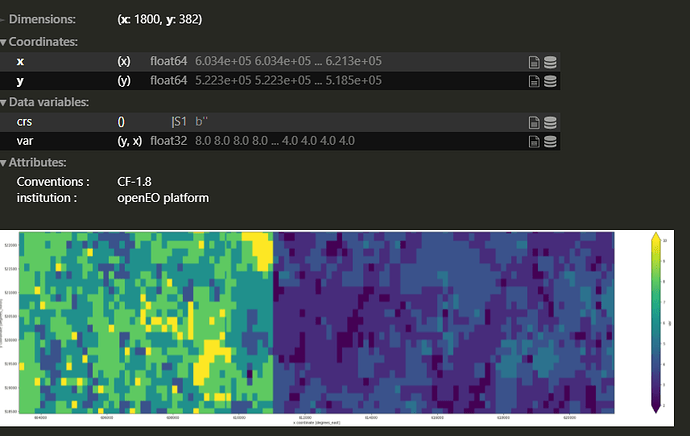
The same ouput with the different scale: