Hi all,
I am creating an MNDWI cube using composite images using a year’s worth of data. Then I am taking the minimum: mndwi_min: DataCube = mndwi.reduce_dimension(dimension="t", reducer="min")
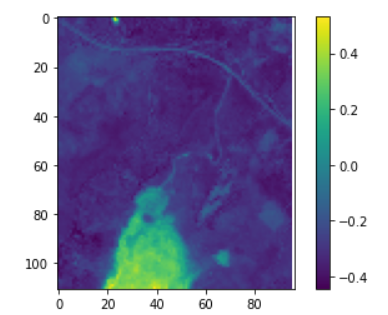
Afterwards I want to threshold this image: mask: DataCube = mndwi_min.apply(lambda val: processes.lte(x=val, y=0.5))
The result is not as expected:
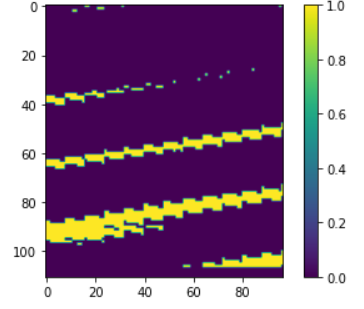
Can you see if I made a mistake somewhere?
process graph of the mask:
{'loadcollection1': {'process_id': 'load_collection',
'arguments': {'bands': ['B11', 'B08', 'B03'],
'id': 'SENTINEL2_L1C_SENTINELHUB',
'spatial_extent': {'west': -5.707740783691406,
'east': -5.6764984130859375,
'south': 36.40221798067486,
'north': 36.4315034892636,
'crs': 'EPSG:4326'},
'temporal_extent': ['2019-01-01', '2022-01-01']}},
'adddimension1': {'process_id': 'add_dimension',
'arguments': {'data': {'from_node': 'loadcollection1'},
'label': 'SENTINEL2_L1C_SENTINELHUB',
'name': 'source_name',
'type': 'other'}},
'renamelabels1': {'process_id': 'rename_labels',
'arguments': {'data': {'from_node': 'adddimension1'},
'dimension': 'bands',
'source': ['B11', 'B08', 'B03'],
'target': ['swir', 'nir', 'green']}},
'resamplespatial1': {'process_id': 'resample_spatial',
'arguments': {'align': 'upper-left',
'data': {'from_node': 'renamelabels1'},
'method': 'cubic',
'projection': None,
'resolution': 30.0}},
'aggregatetemporal1': {'process_id': 'aggregate_temporal',
'arguments': {'data': {'from_node': 'resamplespatial1'},
'intervals': [['2019-01-01 00:00:00', '2020-01-01 00:00:00'],
['2020-01-01 00:00:00', '2021-01-01 00:00:00'],
['2021-01-01 00:00:00', '2022-01-01 00:00:00']],
'labels': ['2019-01-01 00:00:00',
'2020-01-01 00:00:00',
'2021-01-01 00:00:00'],
'reducer': {'process_graph': {'quantiles1': {'process_id': 'quantiles',
'arguments': {'data': {'from_parameter': 'data'},
'probabilities': [0.2]},
'result': True}}}}},
'reducedimension1': {'process_id': 'reduce_dimension',
'arguments': {'data': {'from_node': 'aggregatetemporal1'},
'dimension': 'bands',
'reducer': {'process_graph': {'arrayelement1': {'process_id': 'array_element',
'arguments': {'data': {'from_parameter': 'data'}, 'index': 2}},
'arrayelement2': {'process_id': 'array_element',
'arguments': {'data': {'from_parameter': 'data'}, 'index': 0}},
'subtract1': {'process_id': 'subtract',
'arguments': {'x': {'from_node': 'arrayelement1'},
'y': {'from_node': 'arrayelement2'}}},
'add1': {'process_id': 'add',
'arguments': {'x': {'from_node': 'arrayelement1'},
'y': {'from_node': 'arrayelement2'}}},
'divide1': {'process_id': 'divide',
'arguments': {'x': {'from_node': 'subtract1'},
'y': {'from_node': 'add1'}},
'result': True}}}}},
'reducedimension2': {'process_id': 'reduce_dimension',
'arguments': {'data': {'from_node': 'reducedimension1'},
'dimension': 't',
'reducer': {'process_graph': {'min1': {'process_id': 'min',
'arguments': {'data': {'from_parameter': 'data'}},
'result': True}}}}},
'apply1': {'process_id': 'apply',
'arguments': {'data': {'from_node': 'reducedimension2'},
'process': {'process_graph': {'lte1': {'process_id': 'lte',
'arguments': {'x': {'from_parameter': 'x'}, 'y': 0.5},
'result': True}}}},
'result': True}}